The Accelerator and Maximising Performance on Modern Computers
The Accelerator is able to process hundreds of millions of data rows per second on a single computer using the Python programming language. This post will discuss the performance bottlenecks of a modern computer and explain how the Accelerator is optimized towards maximum speed given these hardware constraints.
The problem
Assume that we have a large dataset stored on disk, and that we want to process it as fast as possible. Fast data processing is important for several reasons, for example
- more data can be processed in the same amount of time,
- shorter test iteration time makes development easier,
- speed can be traded for cheaper hardware,
- a faster solution is more “realtime”.
and so on.
What is Limiting Execution Time?
On a high level, a computer is composed of storage (disk, memory), processing (CPU) hardware, and interconnects there between, see figure below
From a high level perspective, it makes sense to start with these two questions:
- How fast can we read our data from disk?
- How fast can we perform computations on the data?
It is fundamental to understand the bottlenecks of the system. If we understand the bottlenecks, we can take them into account when optimising for speed. This is the key performance design principle for the Accelerator, and in the next sections we’ll take a closer look at how it works and what can be done.
Storage: Random Access Versus Streaming
First, we note that the disk accessing pattern matters. There is a significant average access time associated with accessing a random data block on a standard rotating hard drive. This is because we have to wait for both the disks to rotate and the heads to move to the position where the data we need is stored. Once the data is in position, streaming it from disk to memory is fast.
Hard disk access time is in the range of 10 milliseconds. This implies that we can access about 100 random data chunks per second and drive. Clearly, this is not much.
Access times are significantly improved using solid state storage technology, but at a significantly higher cost.
Access Times Matters
One way to store the data to be analysed is to use a traditional relational database. Such databases are designed for random access. The database does not have a clue about what your next operation will be. Therefore, a lot of effort has been put in to predict, cache, partition, and by other means minimize the average access time per database record.
If we just access a small random set of our data once in a while, 10 milliseconds might not matter and databases are fine. But if we are to process a large chunk of the data, the random access approach will be very slow. In order to speed it up, significant effort has to be spent tuning the database for the particular dataset and application. Even then, throughput will be far from its theoretical optimum. We believe that it is better to work on the actual problem than to spend time and resources tuning particular software systems.
If we are to process a large portion of the dataset, it is much more efficient to stream all or most of the data and filter out and work on the relevant parts only. If streaming is, say 1000 times faster than random access, streaming is better if we process as little as 0.1% of the data or more!
Disk Throughput: How Fast can we Read from Disk?
The next thing to look at is how fast we can stream data from disk to memory. The minimalistic way to read data very fast from disk is like this:
cat file > /dev/null
This command will, as fast as possible and with minimal overhead, read data from disk and discard it. There is no faster way to read data from disk, at least when using a standard file system. If the disk and CPU are fast enough, this will saturate the disk to memory bandwidth.
On a “standard” computer, we reach a few 100MB/s throughput.
Maximizing Throughput
With the disk bandwidth saturated, we can transfer more useful information per second by compressing the data on disk:
zcat file.gz > /dev/null
Decompression is a lightweight operation on a modern CPU. The compression typically increases disk bandwidth by 5 to 10 times on a realistic dataset, so this is a favorable trade-off. (In a data science context, uncompressable datasets are often not that interesting since they contain a minimum amount of patterns and repetitions.)
When storing data in a compressed format, we can stream in the ballpark of 1000MB/s from disk to CPU, which is faster than the disk interface.
Note also that streaming data in a linear fashion from disk to memory and memory to CPU yields a completely predictable addressing pattern. A modern CPU can use this and do cache memory read-ahead that could hide memory accessing times.
Can we go Faster?
Well, as soon as we start to process the data, the CPU load will go up, and we realize that processing speed of the CPU will be the new bottleneck. We have turned a data-intensive problem into a processing-intensive one.
Fortunately, modern computers come with several CPU cores, so we can read and decompress multiple files in parallel, thereby increasing the computational power per data item while maintaining the high disk to memory bandwidth:
zcat file1.gz | processing_program &
zcat file2.gz | processing_program &
...
RAM is Faster than Disk
It makes sense to have a computer with lots of RAM in it. Operating systems such as Linux or FreeBSD use unallocated RAM for disk buffers / cache, which means that when a file is read from or written to disk, it also stays in RAM to allow for much faster access should it be accessed again.
As for disks, RAM does not get much faster over time, but it gets cheaper, i.e. you get more for the same price. It is common, but unnecessarily old-fashioned to use machines with little RAM. For example, AWS EC2 instances with 4TB RAM and even 12TB RAM(!). have been available for a while now. In general, more RAM is better, but even a small increase in RAM may improve performance.
The nice thing is that more RAM may increase performance without any extra work (due to the disk buffering). In addition, data intensive algorithms such as collaborative filtering execute much much faster if the source data fits into RAM.
The Accelerator Approach
The important tricks that we’ve just seen are used by the Accelerator to maximize data read and write performance, in particular:
- data is stored in a compressed format, and
- data is split into several files, slices, and processed in parallel.
In addition, the Accelerator
- splits data into one file per column and slice,
- uses an optimised disk storage format, and
- spreads data into independent slices using a hashing algorithm.
One File Per Column
A dataset may typically have several columns containing different information. Each processing/analysis task typically uses only a few of them in each processing task. Therefore it makes sense to separate the columns into independent files, so that the read bandwidth is occupied only with data that is necessary for the processing task at hand.
The image above shows how a five-column tabular dataset is separated into four slices having five column files each. Column four is marked in light gray.
Optimised Storage Format
Data can be stored very efficiently on disk. If we know that a data column only contains, say, 64 bit floats, we can write that information in the header of the file, and then concatenate all the floats together without separation. Variable length types prefix each data item with a record containing its length. Similar efficient approaches exists for all the various data types available, optimising both storage requirements and parsing times.
In addition, if a dataset is small, slice files are concatenated into fewer larger files, and in-file indexing to access the data. This is done in order to keep the number of small files and total disk access time to a minimum.
Data Partitioning using a Hash Function
Using a hash function to determine which data row that goes into which slice is a simple way to pave the way for entirely independent parallel processing.
The figure below illustrates a small dataset containing pairs of users
and movies. The upper part of the figure shows the original dataset,
while the bottom part illustrates the dataset partitioned into three
independent slices based on a hash function operating on the user
column.
After hash partitioning, any user will appear in exactly one slice. If we do operations per user, such as for example collecting the set of movies for each user, there is no need to merge data between slices. Computations can be carried out in parallel without any communication between processes.
This data partitioning before processing is kind of the opposite of merge after processing that is being used by Map-Reduce. Since hash partitioning is done before data computations, it is carried out only once, even if we do repeated computations. For Map-Reduce, on the other hand, reduction is a necessary step in the processing. Besides, hash partitioning a dataset is a relatively fast operation.
Some Performance Figures
The fastest machine we’ve used produced these numbers in 2017 on an example file with one billion lines and six columns, in total 79GB in size:
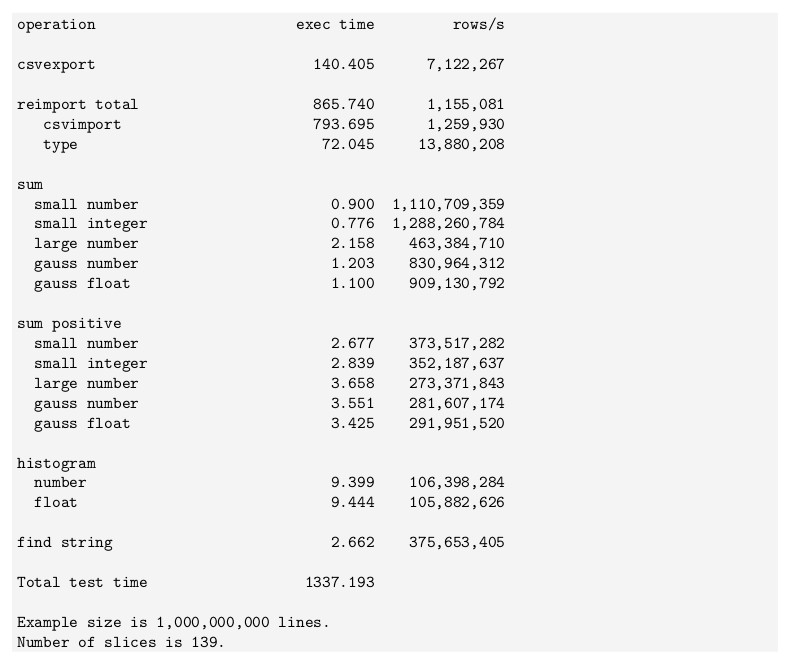
This data is cut from an old version of the
Accelerator’s installation manual, and the
corresponding code is open sourced by eBay. Note that all numbers are
produced using high level Python programs, and that the csvimport
method has been improved significantly in later versions of the
Accelerator.
Summary
The Accelerator is using a set of low level basic techniques such as compression, slicing, partitioning, and parallel execution to mitigate the computer’s bottlenecks and maximize performance. The result is a powerful piece of software that can do incredible things at high speed and very low cost.
Additional Resources
The Accelerator’s Homepage (exax.org)
The Accelerator on Github/exaxorg
The Accelerator on PyPI
Reference Manual